|
Clyde
Hutchison, III, Component Project Leader
Research
BSGC
Role
Clyde A. Hutchison III, will oversee Component Project VII, correlation to cellular
function. He comes to the Center with experience in mycoplasma genomic
analysis, with a focus on identifying the minimal essential gene
set for mycoplamas. He will serve as a liaison between the Center
and the mycoplasma and genomics communities. He will also assemble
the current state of knowledge of function of mycoplasma genes selected
as targets for structural studies at the BSGC.
Component Project
Information
 |
 |
 |
|
| |
|
|
|
|
| |
|
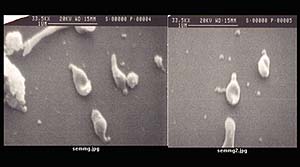
Scanning
electron microscope image of Mycoplasma genitalium.
Photo: K. Frantz, A. Albay and K. Bott, The University of North
Carolina at Chapel Hill. |
|
| |
|
|
|
The Hutchison
lab will provide a liaison between the structural work going on
in the Center, the mycoplasma research community, and the functional
genomics community. In particular we will provide an interface between
the structural work in the Center and ongoing functional genomic
studies of mycoplasmas at The Institute for Genomic Research (TIGR).
We will also undertake experimental approaches to defining the cellular
function of genes selected as candidates for structural studies.
The work under this subproject will be directed toward the following
goals:
1. Collection
of existing information concerning the functions of target genes.
We propose to collect existing information from the literature and
from the mycoplasma research community for each gene selected for
structural studies. If homologs exist in other species we will collect
information on these homologs if the function has not been studied
directly in mycoplasma. The information will be made available in
the form of a spreadsheet over the web for all the participants
in the project.
2. We will extend
our analysis of gene disruptions using transposons to test whether
target genes are essential for growth in the laboratory.
We will use
gene-specific PCR to detect transposon insertion events in target
genes. This could allow the identification of non-essential genes
that have not so far been identified in our published work. We also
propose a strategy using DNA microarrarys to detect gene-specific
transposon disruption events.
3. Detection
of mRNA as a test for expression of the gene.
RT PCR will
be used to make a semiquantitative assessment of the level of expression
of each target gene. As an alternative approach, in collaboration
with TIGR, the expression will be studied using DNA microarray technology.
4. Targeted
gene knockouts as a tool for assessing cellular function.
Since homologous
recombination has been used to specifically disrupt a gene in M. genitalium (MG218), we expect that it can be developed into a tool
that can be used to aid in the determination of unknown cellular
functions.
5. Complementation
of E. coli or yeast mutants.
We will attempt
to complement E. coli or yeast mutants with mycoplasma genes as
a way to identify gene functions.
6. We will
attempt to establish a system for regulated expression of a gene
introduced into mycoplasma.
If this can
be done then the chromosomal copy of the gene could be knocked out
and the phenotypic effect studied. This would permit study of genes
of unknown function that are essential. Establishment of such a
system can use existing methods to introduce a gene using a transposon.
We will attempt to develop a repressor-regulated system based on
the E. coli lac system.
7. Localization
of the gene product to a cellular compartment.
Cells expressing
an epitope tagged mycoplasma gene product will be fractionated and
the distribution of the gene product within the cell will be studied
to obtain clues concerning gene function.
|